Welcome to SpatialCCDB! click thumbnails to quickly browse the spatial cell communication.
Search SpatialCCDB
What is SpatialCCDB?
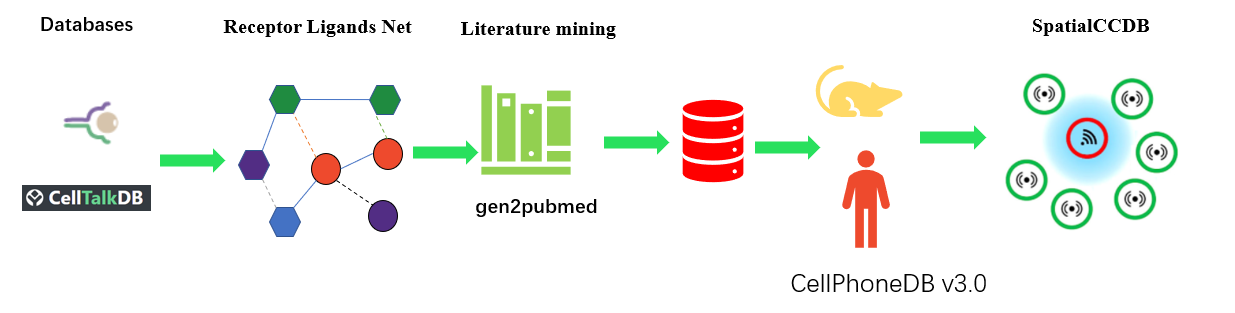
SpatialCCDB is the first public database that specifically curates spatially resolved transcriptomic data from published papers, aiming to provide a comprehensive and accurate resource of spatial cell communication in tissues. Currently, SpatialCCDB contains detailed information of 18 datasets with spatially resolved transcriptome, covering from 12 tissues in two species (Human and Mouse). SpatialCCDB allows users to browse the spatial cell communication of all the 12 tissue online and provides positional information of ligand-receptor relationships. It also provides tissue Receptor-Ligand (LR) genes functional, KEGG and REACT signal pathway, and diseases related to LR genes.
What can SpatialCCDB provide?
1. An atlas of spatially resolved cell communication. The website currently includes datasets of Heart, Large Intestine, Liver, Lung, Ovary, Prostate, Colon, Gastric, Kidney, Skin, Ileum.
2. A web interface for spatial cell communication visualization.
3. An online tool for quick retrieval of spatial cell communication in a certain tissue of interest.
4. Receptor ligand (LR) genes identified by stLearn and CellPhoneDB, enrichment analysis, and diseases related to these LR genes.